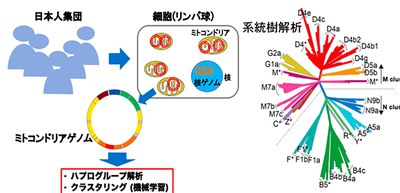
Mitochondrial DNA (mtDNA) specific to Japanese population clarified
Mitochondria, classical membrane-bound organelles of eukaryotic cells, evolve with cells and play an important role in maintaining lives. Mitochondria have their own DNA, or mitochondrial genome, and contain oxidative phosphorylation genes to generate adenosine triphosphate (ATP) to supply readily available energy for the body. A mutation rate in mtDNA is higher than in that of nuclear DNA (nDNA) and mitochondria lack recombination.
Since mitochondrial DNA (mtDNA) is passed from mother to child, testing mtDNA allows for investigation into our maternal line. Maternal haplogroups are determined by assessing mtDNA. Haplogroups are classifications of the mtDNA haplotypes defined according to a set of the specific mtDNA variants.
A research group from Osaka University, by analyzing mtDNA using the whole genome sequence (WGS) of some 2,000 Japanese individuals who participated in the BioBank Japan project, identified the mitochondrial genome sequence specific to the Japanese population, clarifying the difference in mtDNA variant patterns between mtDNA and nuclear nDNA.
Using a set of machine learning methods, the researchers conducted unsupervised clustering of the mtDNA variant patterns to assess their correspondence with defined haplogroups, visualizing haplogroup diversity within the Japanese population. Their haplogroup analysis demonstrated that frequency spectra of the haplogroups were population-specific and were heterogeneous among geographical regions within Japan.
Together with WGS-based mtDNA variant imputation*1, the researchers conducted a phenome-wide association study of 147,437 Japanese individuals with 99 clinical phenotypes (46 complex diseases, 49 quantitative traits, and 4 drinking and smoking habits) in order to identify mtDNA variants associated with human complex traits. As a result, they clarified associations of the mtDNA variant with creatine kinase (an enzyme highly expressed in tissues that rapidly consume energy such as skeletal muscle and brain), eGFR (estimated glomerular filtration rate showing the level of renal function), AST (aspartate aminotransferase, a liver function-related trait), and Graves’ disease (an immune-related disease).
This group revealed Japanese population-specific mitochondrial genome sequences, demonstrating that the frequency distribution of each haplogroup were heterogeneous among geographical regions within the country. The clarification of population-specific mtDNA structure and their pleiotropic associations with human complex traits will accelerate the understanding of the association between mitochondrial genomes and diseases.
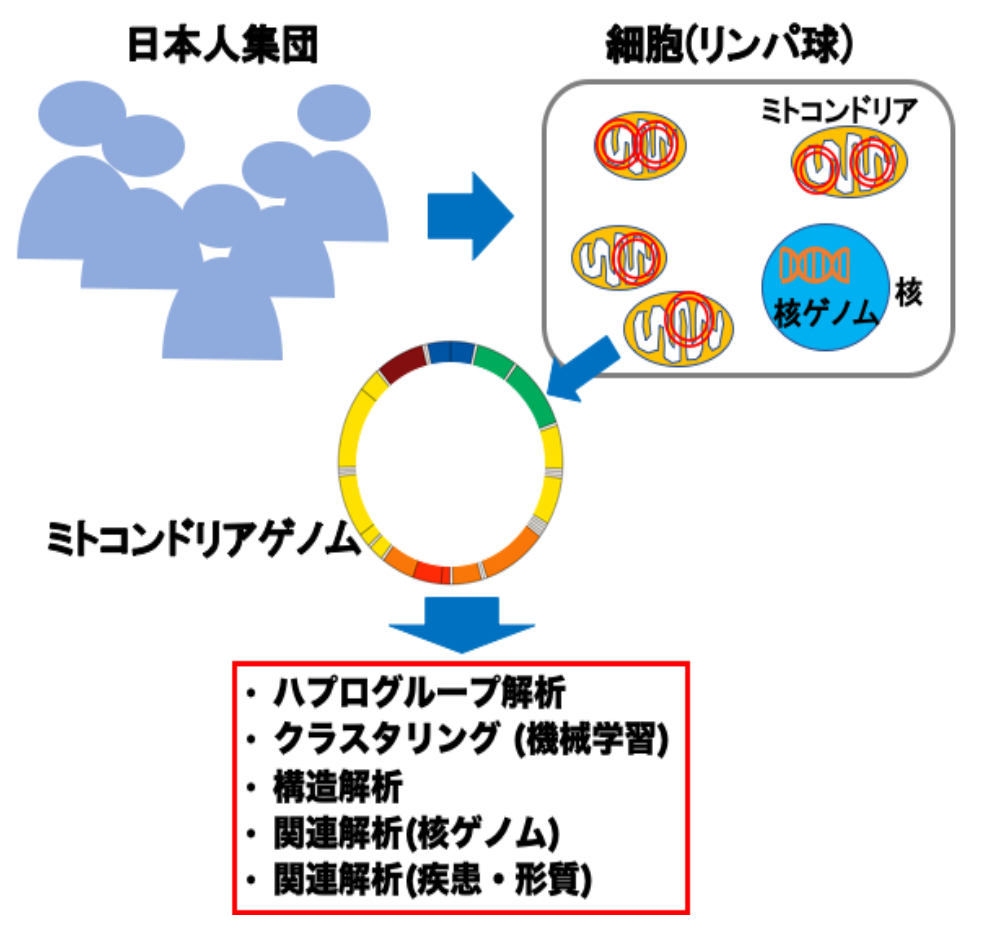
Figure 1
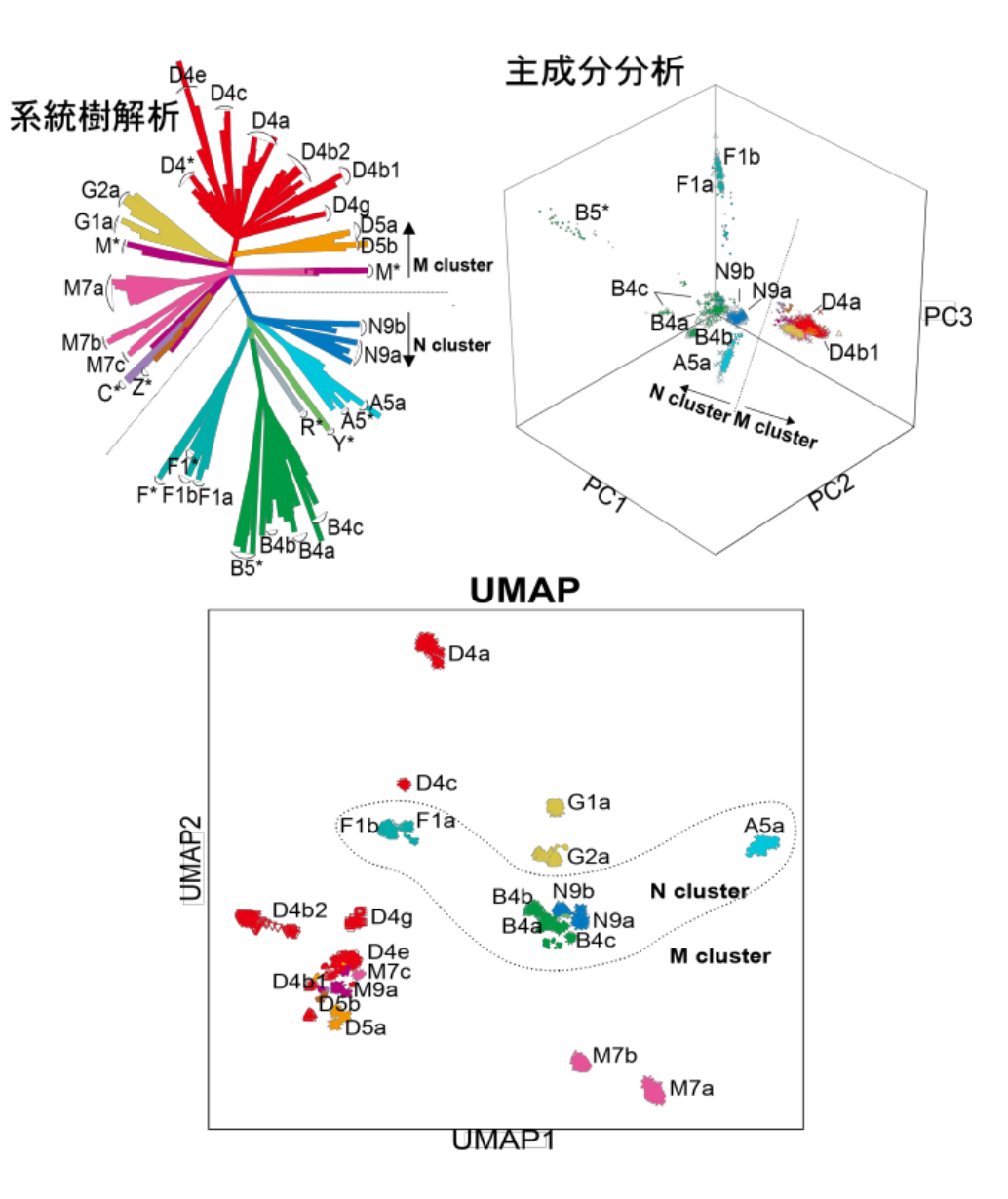
Figure 2
The article, “Genetic and phenotypic landscape of the mitochondrial genome in the Japanese population” was published in Communications Biology at DOI: https://doi.org/10.1038/s42003-020-0812-9.
