The discovery of a totally new kind of “mark” in human cell nucleus
Will lead to the clarification of mechanism of development of metabolism and cardiovascular disorder
A group of researchers led by Professor ISAKA Yoshitaka (Division of Medicine, Graduate School of Medicine, Osaka University), Endowed Chair Professor TAKAHARA Shiro (Department of Advanced Technology for Transplantation, Graduate School of Medicine, Osaka University), Endowed Chair Professor KAIMORI Junya and Professor TAKAO Toshifumi (Institute for Protein Research, Osaka University), Professor KIMURA Hiroshi (Graduate School of Bioscience and Biotechnology, Tokyo Institute of Technology), and Professor OKAWA Yasuyuki (Medical Institute of Bioregulation, Kyushu University), verified the presence of Histone H4 lysine 20 acetylation (H4K20ac) by using mass spectrometry. H4K20ac is a mark of the cell nucleus but whose existence in mammalian cells has been indirectly proven.
This group made antibodies against H4K20ac. Although known acetylization was enriched around promoters of highly expressed genes, this group, by using a chromatin immunoprecipitation method, next-generation sequencing, and supercomputers, found that H4K20ac, a new type of histone modification, was enriched around promoters of minimally expressed genes.
In recent years, it has been known that epigenetic factors, factors outside of the genetic sequence, are deeply involved in the development of various diseases. Histone, a component of chromatin consisting of DNA, receives post-translational modification for changing the function of protein as an epigenetic factor.
It is expected that the mechanism of development of various diseases such as metabolic disorders including diabetes, cardiovascular diseases including cardiomegaly, kidney disorders, and cancer will be elucidated by using H4K20ac, a new type of histone modification discovered by this group.
Author summary
Epigenetics is a research field where not DNA sequence but DNA methylation or histone acetylation are studied as a tool for conveying traits to the next generation. In these days, epigenetics is reported to be implicated with pathogenesis or disease progression. So it has attracted lots of attention. Histone acetylation is generally associated with gene activation. We identified H4K20ac, which was discovered only in plant cells. To understand the function of H4 lysine 20 acetylation (H4K20ac), we have developed a specific monoclonal antibody and performed ChIP-seq analysis using HeLa-S3 cells. H4K20ac was enriched around the transcription start sites (TSSs) of minimally expressed genes and in the gene body of expressed genes, in contrast to most histone acetylation being enriched around the TSSs of expressed genes. The distribution of H4K20ac showed little correlation with known histone modifications, including histone H3 methylations. A motif search in H4K20ac-enriched sequences, together with transcription factor binding profiles based on ENCODE ChIP-seq data, revealed that most transcription activators are excluded from H4K20ac-enriched genes and a transcription repressor NRSF/REST co-localized with H4K20ac. These results suggest that H4K20ac is a unique acetylation mark associated with gene repression. So our discovery of a new kind of histone acetylation modification, H4K20ac will widen the field of view in epigenetics. It is highly expected that using this new histone modification, the new knowledge in pathogenesis or disease progression will be clarified in the future study.
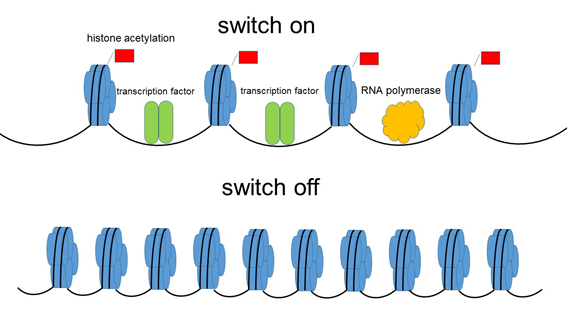
Figure 1. It was accepted that histone acetylation modifications are implicated only in gene activation,switch on”.
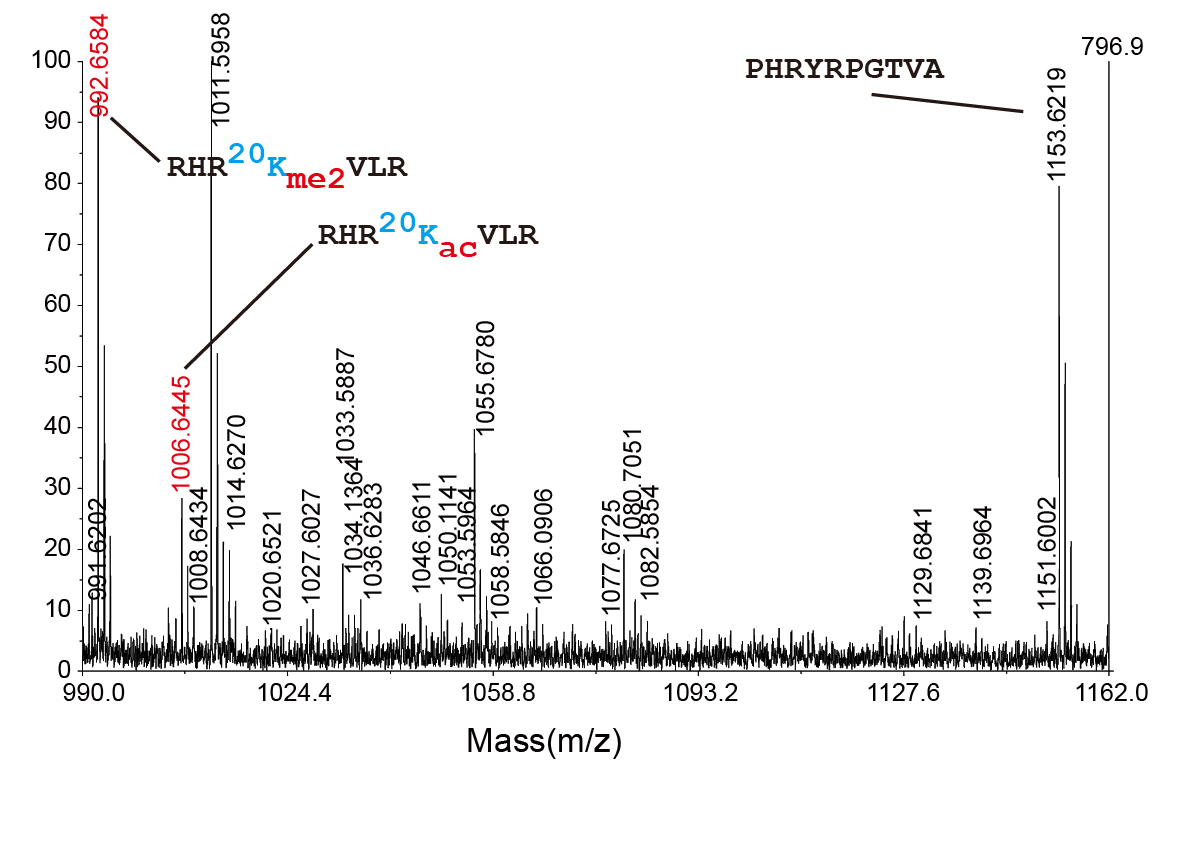
Figure 2. We confirmed the existence of H4K20ac in human cells by measuring accurate mass of the peptide with K20ac.
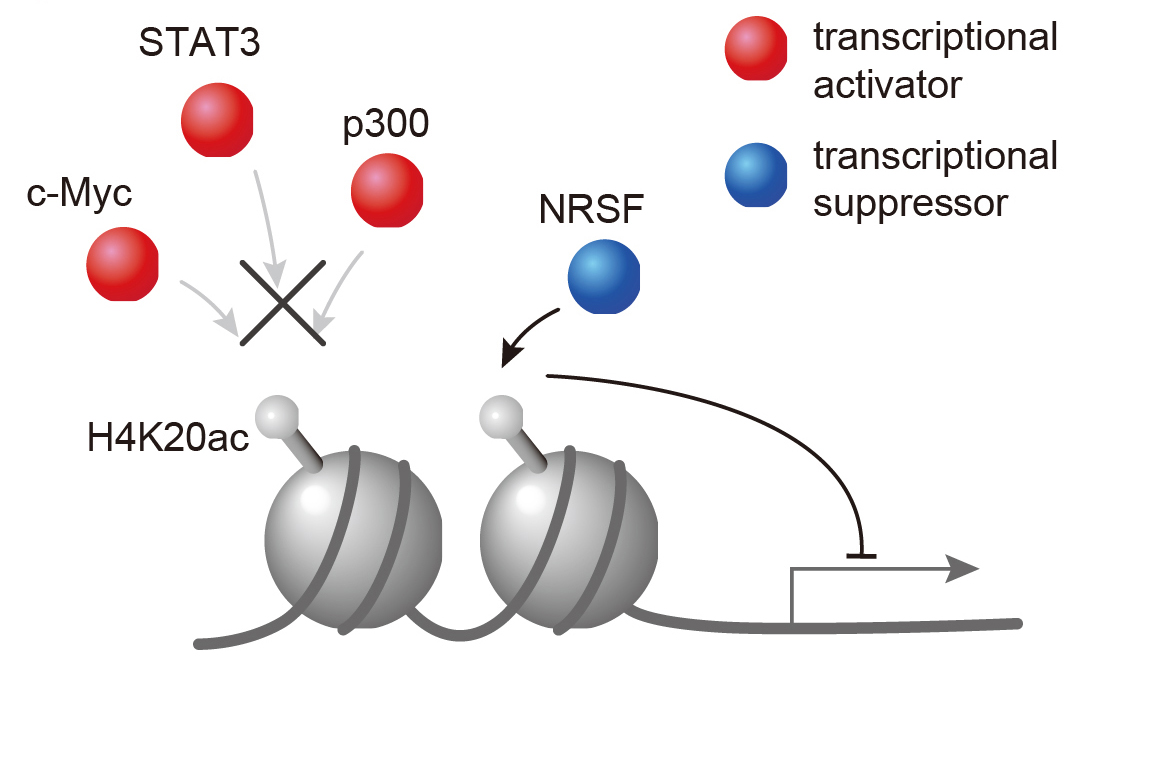
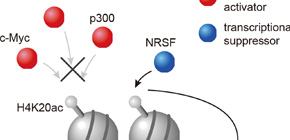
Figure 3. H4K20ac accumulated region excludes transcriptional activators but includes only a transcriptional suppressor. Therefore, H4K20ac is implicated in gene repression.
To learn more about this research, please view the full research report entitled " Histone H4 lysine 20 acetylation is associated with gene repression in human cells " at this page of the Nature website.
Related link
