Clarified mechanism of rotation of node cilia-principal for asymmetry of the body
Division of Biotechnology and Life Science, Institute of Engineering,Tokyo University of Agriculture and Technology -- SHINOHARA Kyosuke , Specially Appointed Associate Professor
Graduate School of Frontier Biosciences, Osaka University -- HAMADA Hiroshi , Professor
Our bodies look symmetrical, but most internal organs are asymmetric in shape or in position. In mouse embryos, a model animal that is closest to humans, cell groups, which are a source of organs, are symmetrical, but become asymmetric some 8 days after fertilization.
A research group of Osaka University and Tokyo University of Agriculture and Technology, together with collaborative research institutes, clarified the mechanism of rotation of node cilia which determines the left-right asymmetry of the body and elucidated part of the relationship between the ciliary structure and ciliary motility, which had little experimental knowledge. This group's achievement may lead to the clarification of causes of heterotaxia, bronchitis, and infertility caused by impaired motility of cilia and flagella.
In mice, a model animal closest to humans, the body's left and right are determined 8 days after fertilization. It is thought that at this time, cilia of some 200 cell groups called 'nodes' which appear on the midline of the body (a 2-5μm motile protuberance that project from the cell body) rotate clockwise and generate a leftward fluid flow (nodal flow), which breaks the bilateral symmetry of genes that are expressed on the left and right of the node and determines left and right in the body.
By this time, the anterior-posterior polarity in mouse embryos has been established. The leftward flow is generated by a combination of two features of node cilia: posterior tilt and their clockwise rotation.
Regarding how node determines the anterior-posterior polarity of a cell, the involvement of PCP (Planar Cell Polarity) signaling positions has been reported in recent years; however, the mechanism of ciliary clockwise rotation had been unknown.
In cooperation with Beijing Institute of Technology, Research Center for Ultra-High Voltage Electron Microscopy, Osaka University, and RIKEN Center for Life Science Technologies, Kyosuke Shinohara (Specially Appointed Associate Professor, Division of Biotechnology and Life Science, Institute of Engineering, Tokyo University of Agriculture and Technology) and Hiroshi Hamada (Professor, Graduate School of Frontier Biosciences, Osaka University) found treatment with Taxol, an anticancer drug that stabilizes microtubules, disturbed the motility pattern of node cilia.
Normally microtubules of node cilia are regularly arranged near the membrane. This group examined them with an electron microscope and found that the regular arrangement of microtubules was disturbed in Taxol-treated node cilia.
This group performed computer simulation of ciliary motility based on the electron tomography data in order to clarify a relation between the structure and ciliary motion pattern and found that when the arrangement of microtubules became abnormal, the motion pattern was also disturbed.
From these findings, it was found that in order for node cilia to make a rotation in one direction, regular arrangement of microtubules was necessary. (Figure 1)
Mice possess two types of motile cilia: a node cilium without a central pair and a motile cilium with central-pair microtubules and radial spokes. Motile cilia are found in the airway, oviduct, and brain ventricles.
This group made a hypothesis that Taxol treatment changed the structure of node cilia because they had no central structure for supporting microtubule arrangement and verified the hypothesis.
First, the group treated airway cilia of wild mice with Taxol, but no changes were found in their ciliary motion and structure. Then, the group created mice lacking the radial spoke head protein Rsph4a. The group observed the airway ciliary motion. In wild-type mice, they showed planar motion, but airway cilia in the knockout mice showed clockwise rotation as node cilia did. Finally, when this group treated the airway cilia of the knockout mice with Taxol, the arrangement of microtubules was disorganized due to Taxol exposure and clockwise rotation changed to random rotation.
From these findings, it was suggested that radial spokes played a role of supporting regular arrangement of microtubules by connecting surrounding microtubules with the central structure.
Node cilia need to make L-R polarity based on information on the anterior-posterior polarity in mouse embryos. For that purpose, rotation, not planar motion, is necessary. It is thought that this is the reason why the central structure was lost during the course of evolution.
This research was featured in Developmental Cell (US) on October 26, 2015.
Abstract
Determination of left-right asymmetry in mouse embryos is established by a leftward fluid flow that is generated by clockwise rotation of node cilia. How node cilia achieve stable unidirectional rotation has remained unknown, however. Here we show that brief exposure to the microtubule-stabilizing drug paclitaxel (Taxol) induces randomly directed rotation and changes the ultrastructure of node cilia. In vivo observations and a computer simulation revealed that a regular 9+0 arrangement of doublet microtubules is essential for stable unidirectional rotation of node cilia. The 9+2 motile cilia of the airway, which manifest planar beating, are resistant to Taxol treatment. However, the airway cilia of mice lacking the radial spoke head protein Rsph4a undergo rotational movement instead of planar beating, are prone to microtubule rearrangement, and are sensitive to Taxol. Our results suggest that the absence of radial spokes allows node cilia to rotate unidirectionally but, as a trade-off, renders them ultrastructurally fragile.
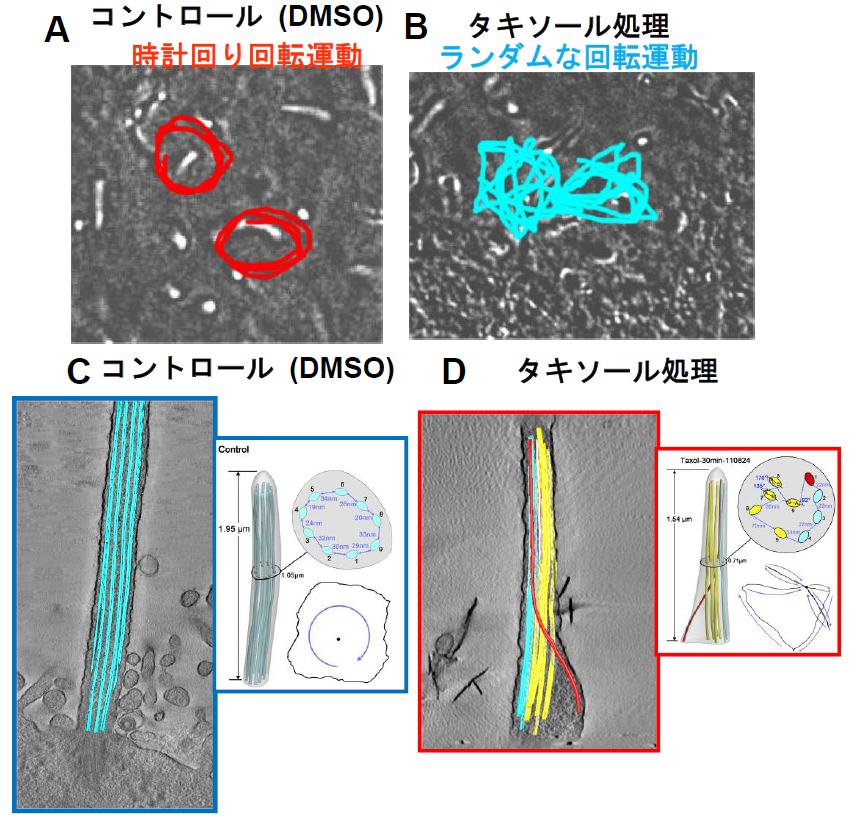
Figure 1
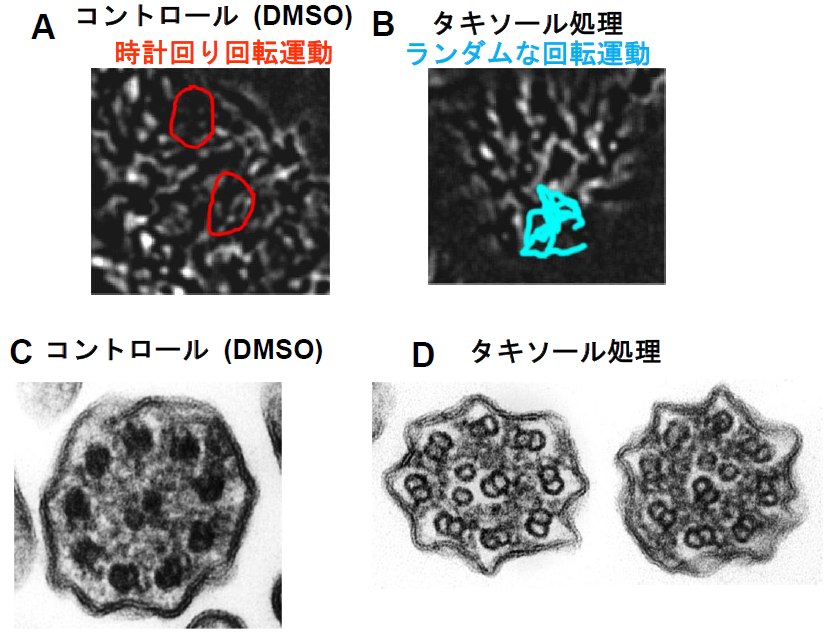
Figure 2
To learn more about this research, please view the full research report entitled “ Absence of Radial Spokes in Mouse Node Cilia Is Required for Rotational Movement but Confers Ultrastructural Instability as a Trade-Off ” at this page of the Developmental Cell website.
Related links
