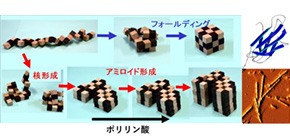
Uncovering Possible Role of Polyphosphate in Dialysis-Related Amyloidosis
Researchers from Osaka University find that a naturally occurring biopolymer encourages the formation of rogue protein aggregates that can lead to dialysis-related disease
Long-term dialysis treatment is tough on the body in many ways, but one of the most serious complications is dialysis-related amyloidosis, a disease characterized by abnormal buildup of protein aggregates—called amyloid fibrils—in joints, tissues and organs.
Although there is no cure, recent research by Osaka University researchers sheds light on how amyloid fibrils form and aggregate, thereby allowing the identification of new therapeutic and preventive avenues to further improve the lives of sufferers.
In a study published in Proceedings of the National Academy of Sciences of the United States of America , Osaka University researchers have identified that polyphosphate (polyP), which is a naturally occurring polymer in the body and a food and drink additive, induces the formation of amyloid fibrils from β2 microglobulin proteins.
PolyPs play an important role in life as storage of phosphorous. When kidney function is normal, β2 microglobulin serum blood levels remain low. However, β2 microglobulin are not efficiently removed from the body in advanced stages of kidney disease. The implementation of dialysis further accentuates this problem, because β2 microglobulin is inable to pass through the dialysis membrane to be filtered.
“Because blood concentration of β2 microglobulin is higher when dialysis is used, the increased level of β2 microglobulin is the most important risk factor. However, the detailed mechanism triggering amyloid formation remains unclear. We wanted to investigate the effects of polyPs, compounds recently reported to be effective for other amyloid proteins, on the formation of amyloid fibrils from β2 microglobulin in an effort to learn how we may prevent it,” says Dr. Yuji Goto.
To do this, the researchers monitored the effect of polyP on the formation of amyloid fibrils from β2 microglobulin under acid and neutral conditions. Under acidic conditions, low polyP concentrations resulted in protein aggregates with an amyiloid-fibril structure but high polyP concentrations induced unstructured (or amorphous) aggregates.
“The results observed under acidic conditions were characteristic of a competitive mechanism,” explains Chun-ming Zhang, previous graduate student of the Institute for Protein Research at Osaka University. “Based on this, we hypothesized that the tendency to form amyloid fibrils or amorphous aggregates is dependent upon the chemical environment.”
However, under normal pH conditions, the results were quite different. The amyloid fibrils that formed were thicker; polyPs promoted amyloid fibril formation through protein aggregation by salting-out effects.
This is the first evidence resolving how polyPs affect amyloid formation at different pH. These findings have clinical application potential because monitoring polyP levels in people receiving long-term hemodialysis could provide information regarding an individual’s risk of dialysis-related amyloidosis.
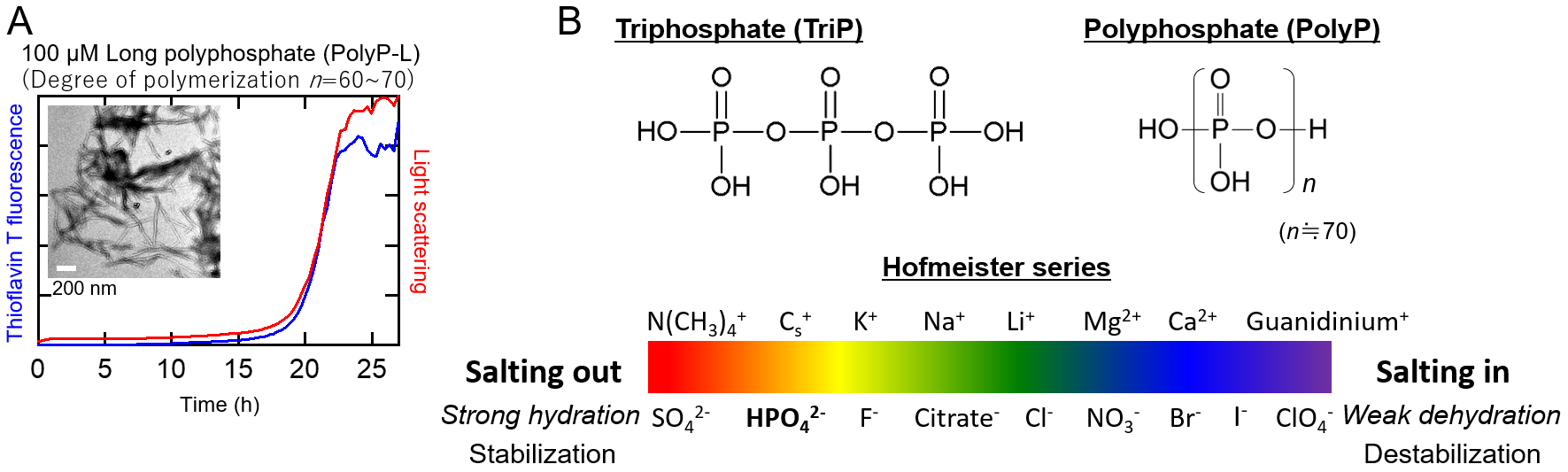
Fig.1 (A) Long polyphosphate (PolyP-L)-induced amyloid formation of b 2 -microglobulin at neutral pH and TEM image. (B) Chemical structures of triphosphate (triP) and polyphosphate (polyP) (top) and the Hofmeister series (bottom).
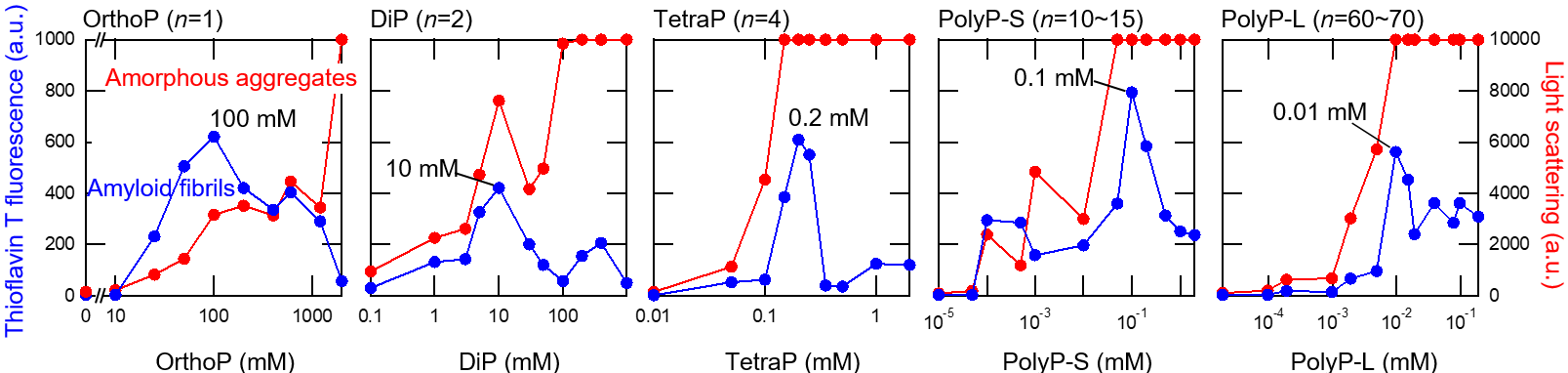
Fig.2 Amyloid formation in the presence of various chain lengths of polyPs at acid pH. Amyloid formation was monitored by thioflavin T fluorescence (blue) and total amount of aggregate formation (amyloid fibrils and amorphous aggregates) was monitored by light scattering (red). The optimum polyPs concentrations for amyloid formation decreased with an increase of chain length of polyPs.

Fig.3 Possible mechanisms of polyP-induced amyloid fibril formation at the acidic and neutral pH conditions.
The article, “Possible mechanisms of polyphosphate-induced amyloid fibril formation of β2-microglobulin,” was published in Proceedings of the National Academy of Sciences of the United States of America at DOI: doi.org/10.1073/pnas.1819813116 .
Related links
