New method reliably supplies protein microcrystals for serial femtosecond crystallography
Determination of 3D crystal structures possible with small samples
A group of researchers affiliated with RIKEN, Osaka University, Kyoto University, and Japan Synchrotron Radiation Research Institute (JASRI) succeeded in developing a generally applicable method for supplying protein crystals for serial femtosecond crystallography making use of the X-ray laser (SFX) at the X-ray Free Electron Laser (XFEL) Facilities SACLA.
• SPring-8 Center, Riken -- IWATA So , Group Director (Professor, Graduate School of Medicine, Kyoto University), SUGAHARA Michihiro , Senior Scientist, Bio-specimen platform Group; NANGO Eriko , Senior Scientist, SACLA Science Research Group
• Osaka University -- MIZOHATA Eiichi , Assistant Professor, Graduate School of Engineering; SUZUKI Mamoru , Associate Professor, Institute for Protein Research
• Kyoto University -- MASUDA Tetsuya , Assistant Professor, Graduate School of Agriculture; SHIMAMURA Tatsuro , Project Lecturer, Graduate School of Medicine; Pan Dong-Qing , former Researcher, Graduate School of Pharmaceutical Sciences
• Japan Synchrotron Radiation Research Institute (JASRI) -- TONO Kensuke , Associate Senior Researcher, JASRI XFEL Research Promotion Room
In order to determine the 3D crystal structure of proteins by atomic-resolution structural investigation, X-ray crystallography making use of protein crystals is suitable. When using radiation light at the synchrotron radiation facility SPring-8 for that purpose, generally, protein crystals of about 30 µm are needed. However, obtaining protein crystals of 30 µm or bigger, especially, obtaining a large enough volume of proteins derived from animals, as well as humans, for research applications in drug discovery making use of crystallization is difficult. Deposited crystals don't grow big enough for use in diffraction experiments. Damage to protein crystal from radiation during experiments is also a big problem.
If X-ray crystal structure analysis using SFX at SACLA can be achieved, minute structures --nanometer- to micrometer-sized protein crystals-- can be used without X-ray radiation damage. Furthermore, the response process in a femtosecond to picosecond when a series of structural change associated with enzymic reaction can be observed with SFX.
However, in order to continuously supply protein crystals to the irradiation point of X-ray laser, it has been necessary to inject liquid samples at a high speed, which means a lot of samples are needed. Adding to this problem, depending on the composition of samples, salt crystals may deposit during the experiment. This is why a stable supply of protein crystals has been difficult to obtain for experiments.
This joint research group succeeded in developing a new technique for conducting diffraction experiments with a variety of proteins by mixing protein crystals with viscous grease and then extruding samples at a low speed. The necessary sample volume for this technique is less than 1 mg, a tenth to a hundredth of that necessary using conventional methods. This technique enabled them to determine 3D crystal structures with small samples of proteins.This group's technique can be used in a variety of research fields in organic and inorganic matter as well as for proteins.
Abstract
Serial femtosecond X-ray crystallography (SFX) has revolutionized atomic-resolution structural investigation by expanding applicability to micrometer-sized protein crystals, even at room temperature, and by enabling dynamics studies. However, reliable crystal-carrying media for SFX are lacking. Here we introduce a grease-matrix carrier for protein microcrystals and obtain the structures of lysozyme, glucose isomerase, thaumatin and fatty acid–binding protein type 3 under ambient conditions at a resolution of or finer than 2 Å.

Figure 1
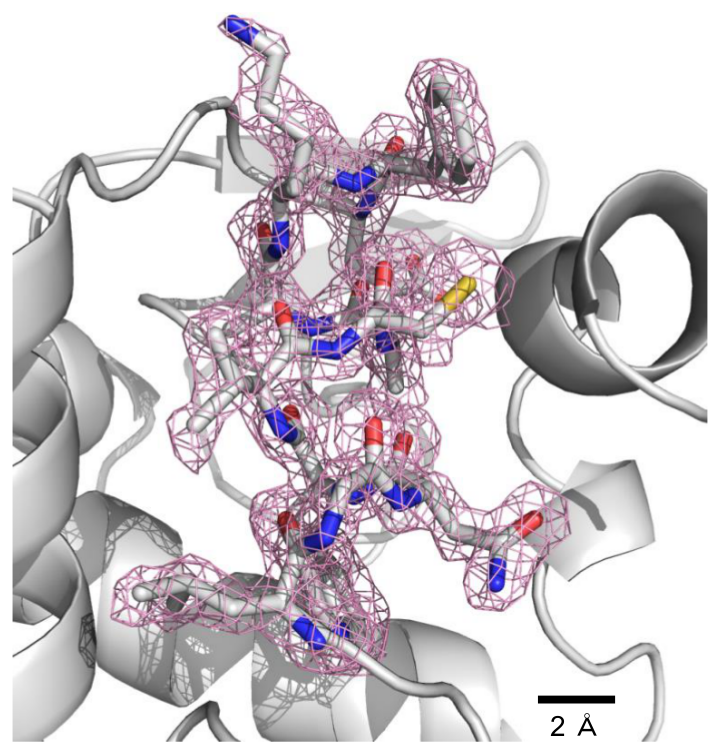
Figure 2
To learn more about this research, please view the full research report entitled " Grease matrix as a versatile carrier of proteins for serial crystallography " at this page of the Nature Methods website.
Related links :
