A gut feeling: microbiome changes may mean early detection of colorectal cancer
Researchers reveal specific changes in the gut microbiota at different stages of colorectal cancer that may lead to important detection methods or further understanding of colorectal cancer development
The gut has a population of organisms that live within in it, called the gut microbiome, which are linked to human health and disease. Recent studies have shown that assessing the genetic changes in fecal samples can accurately reflect the status of the gut microbiome, and may be useful for the early diagnosis of diseases.
A group of researchers from Osaka University have recently reported increases in specific microbiome organisms that are linked to the malignancies associated with colorectal cancer, such as intramucosal carcinomas and polypoid adenomas. Their results, recently published in Nature Medicine Letters , reveal that these specific markers could help distinguish cases of colorectal cancer from healthy samples.
“We believe that colorectal cancer is fundamentally not only a genetic but also a microbial disease,” says one of the study’s corresponding authors, Shinichi Yachida. “Our results show that changes in the gut microbiome are present at the very early stages of colorectal cancer development, which could potentially provide vital diagnostic and causative clues for this disease.”
Colorectal cancer, the third most prevalent cancer globally, is a relatively slow-moving disease—meaning it takes a long period of time before reaching its final, fatal stages. Therefore, early detection is crucial to ensuring effective treatment. The researchers used fecal samples from a little over 600 patients who underwent colonoscopy to assess the characteristics of their gut microbiota and how they relate to colorectal cancer.
“Our results revealed that colorectal cancer was linked to an increase in certain factors in the gut microbiome, as well as the presence of cancer-associated organisms,” says the second corresponding author, Takuji Yamada. “Future studies will focus on the relationship between the gut microbiome and tumor characteristics in individual patients with colorectal cancer. This will help us understand the roles of the microbiome in the development of colorectal cancer.”
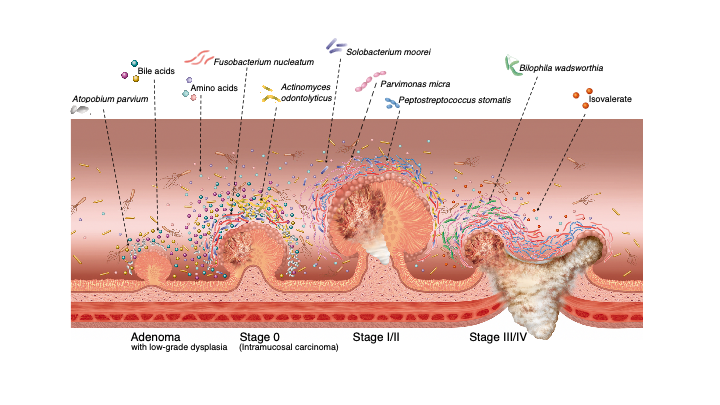
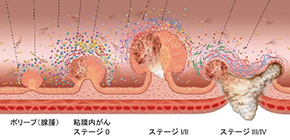
Figure 1. Microbial dynamics during multistep colorectal cancer progression. Graphic representation of major microbial and metabolomic alterations during multistep colorectal cancer progression.
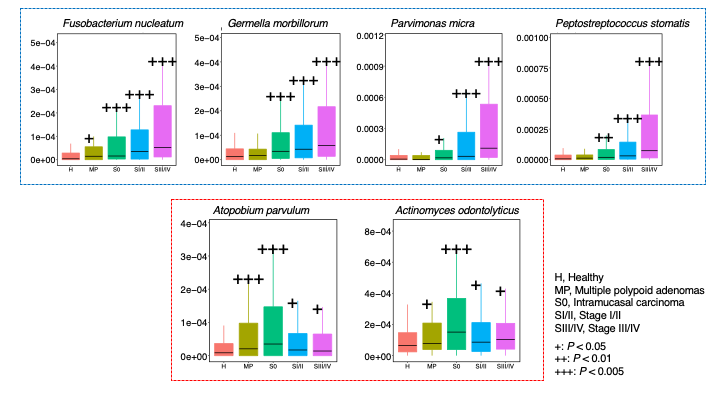
Figure 2. Distinct stage-specific taxonomic signatures with cancer progression. We noted two patterns of significant (P<0.005) species elevation: one that increased across early to later stages ( blue dotted box ), and the other elevated only in the early stages ( red dotted box ). The former was characterized by Fusobacterium nucleatum , Gemella morbillorum , Parvimonas micra and Peptostreptococcus stomatis . The latter pattern was characterized by Atopobium parvulum and Actinomyces odontolyticus .

Figure 3. Metabolome shift in very early stages of tumorigenesis. Compared to the healthy controls, we found significant elevation of deoxycholate (DCA) in multiple polypoid adenomas, and glycocholate and taurocholate in intramucosal carcinomas. Elevation was noted for branched-chain amino acids (isoleucine, leucine, and valine), phenylalanine, tyrosine, serine and glycine. BCFA, branched-chain fatty acid.
The article, “Metagenomic and metabolomic analyses reveal distinct stage-specific phenotypes of the gut microbiota in colorectal cancer” was published in Nature Medicine Letters at DOI: https://doi.org/10.1038/s41591-019-0458-7 .
Related links
