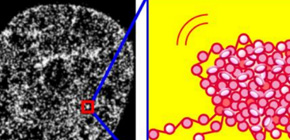
Chromatin structures in live cells revealed
Will clarify genome information and cell-related diseases
It was long believed that DNA is regularly organized in helical folding structures in the nucleus. Recently, however, it has been clarified that these regular structures don’t exist and that DNA is irregularly organized in the nucleus, but no decisive evidence supporting the existence of DNA organized in live cells was found.
A group of researchers led by Researcher NOZAKI Tadasu and Professor MAESHIMA Kazuhiro at the National Institute of Genetics, Research Organization of Information and Systems; Professor NAGAI Takeharu at Osaka University; and Team Leader OKADA Yasushi at RIKEN succeeded in observing DNA organized in live cells by combining super-resolution imaging (photoactivated localization microscopy [PALM]), whose resolution is far better than that of an optical microscope, a world first. As a result, it was found that DNA is irregularly folded and forms compact clutches called ‘chromatin domains.’
Because these chromatin domains are maintained through cell proliferation and cell differentiation, it is suggested that they function as blocks of chromosomes (functional units) for retrieving, reading, and maintaining genome information. This group observed chromatins in live cells by combining super-resolution imaging (PALM), whose resolution is far better than that of an optical microscope and is capable of observing nucleosomes one by one. As a result, this group succeeded not in observing chromatin fibers, but rather, compact clutches (chromatin domains) of about 160 nm in diameter, in which a large number of nucleosomes were irregularly folded, dynamically moving within a single nucleus, a world first.
It was also found that the nature of chromatin domains was produced by two important factors: bonding between nucleosomes and cohesion proteins necessary for bundling chromatins. In addition, these chromatin domains were also found in chromosomes during cell differentiation. From this, it is thought that chromatin domains serve as blocks of chromosomes (functional units) for retrieving, reading, and maintaining of genome information. It is thought that chromatin domains regulate protein accessibility by building blocks and are involved in control and efficiency of information retrieval. It has been also learned that improper DNA folding may cause various abnormalities such as cancerization. It is hoped that this group’s achievements will promote the understanding of abnormality of cells and related diseases.
Abstract
The eukaryotic genome is organized within cells as chromatin. For proper information output, higher-order chromatin structures can be regulated dynamically. How such structures form and behave in various cellular processes remains unclear. Here, by combining super-resolution imaging (photoactivated localization microscopy [PALM]) and single-nucleosome tracking, we developed a nuclear imaging system to visualize the higher-order structures along with their dynamics in live mammalian cells. We demonstrated that nucleosomes form compact domains with a peak diameter of ∼160 nm and move coherently in live cells. The heterochromatin-rich regions showed more domains and less movement. With cell differentiation, the domains became more apparent, with reduced dynamics. Furthermore, various perturbation experiments indicated that they are organized by a combination of factors, including cohesin and nucleosome-nucleosome interactions. Notably, we observed the domains during mitosis, suggesting that they act as building blocks of chromosomes and may serve as information units throughout the cell cycle.
Figure 1
Figure 2
Figure 3
To learn more about this research, please view the full research report entitled " Dynamic organization of chromatin domains revealed by super-resolution live-cell imaging " at this page of the Molecular Cell website.
Related link
