Elucidated the mechanism that determines the zones and initiation of DNA replication in human cell genome
When cells multiply, their genomic DNA is accurately duplicated. However, when abnormalities occur in this mechanism, they can cause cellular aging, cancer, and hereditary diseases. Therefore, the question of how and where DNA replication is initiated in human cells has been a fundamental mystery in life science. An international collaborative research team including Professor Masato Kanemaki of the National Institute of Genetics has developed a new technology, the ligase-depletion Okazaki fragment sequencing (LD-OK-seq) method, that enables highly precise detection of replication initiation zones in the human cell genome and has elucidated the principles behind the initiation of DNA replication. The results of this study revealed that human cells have a flexible mechanism that allows them to initiate replication from almost anywhere in the genome, except within genes that are being transcribed. This is because the MCM helicase required for DNA replication is widely bound throughout the genome. On the other hand, it was found that zones where replication is particularly prone to initiate in early S phase are selected by the binding of a protein called TRESLIN-MTBP to MCM, and that a regulatory mechanism is also at work that antagonistically controls this binding. These findings are the first to elucidate the mechanism by which human cells determine replication initiation zones, paving the way for understanding cancer, aging, and genetic diseases associated with abnormal DNA replication. It is expected that this will lead to the development of technology to artificially control DNA replication in the future. This research was conducted as an international collaboration led by the research group of Professor Masato Kanemaki of the National Institute of Genetics, and also included Project Leader Yasukazu Daigaku of the Japanese Foundation for Cancer Research, Professor Chun-Long Chen of the Institut Curie, France, and Professor Chikashi Obuse of the Graduate School of Science, the University of Osaka.
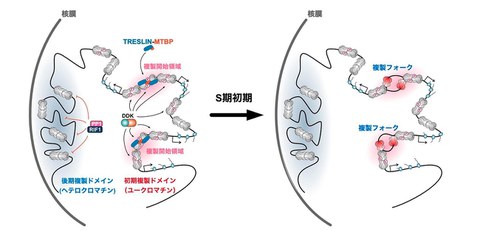
Fig. MCM helicases are widely bound to genomic DNA, and their phosphorylation is antagonistically controlled by the kinase DDK and the RIF1-PP1 de-phosphorylates. Replication initiation zones are determined when TRESLIN–MTBP recognizes the phosphorylated MCM helicase (left). MCM helicase is activated and replication forks are formed (right).
Credit: Chikashi Obuse
Research Background
Until now, research into DNA replication has focused mainly on microorganisms such as E. coli and yeast. In these organisms, the zone where DNA replication initiates (replication origin) is determined by the DNA sequence. However, in many eukaryotic cells, including humans, the DNA sequence at which replication initiates is not determined by the DNA sequence itself, and for many years it remained a mystery as to where replication initiates in the human genome and how that zone is selected. In addition, DNA replication occurs during the S phase of the cell cycle, with euchromatin replicating in the early S phase and heterochromatin replicating in the late S phase. This is called replication timing (RT) and is conserved across species. However, the mechanism that generates RT was unknown.
Research Contents
The research team developed a new technology, ligase-depletion Okazaki fragment sequencing (LD-OK-seq) method, for highly accurate detection of DNA replication initiation zones in the human cell genome. Furthermore, by analyzing the proteins that bind to these zones, the researchers clarified the basic principle of how human cells determine the replication initiation zone.
It is elucidated that human cells have the ability to initiate DNA replication almost anywhere, except in the zone of a transcribed gene. This ability is due to the widespread binding of MCM helicase, an enzyme required for DNA replication, throughout the genome. On the other hand, the research group found that in the early S phase, replication frequently initiates in the zones between transcribing genes (intergenic regions), and that the zone is determined by the binding of a protein called TRESLIN-MTBP, which activates the MCM helicase. Furthermore, they discovered an antagonistic regulatory system that controls the binding of TRESLIN-MTBP to MCM.
In addition, it was revealed that in cells lacking RIF1, which is involved in the antagonistic regulatory system that controls binding to MCM, the binding between MCM helicase and TRESLIN-MTBP becomes randomized, causing DNA replication to occur from anywhere on the genomic DNA. As a result, replication timing was lost.
This study revealed that human cells possess a flexible system in which the potential for replication is widely distributed across the genome, and then the zone of replication initiation is determined by the binding of TRESLIN-MTBP. It is also found that replication timing is controlled via TRESLIN-MTBP.
Fig. MCM helicases are widely bound to genomic DNA, and their phosphorylation is antagonistically controlled by the kinase DDK and the phosphatase RIF1-PP1. When TRESLIN-MTBP recognizes phosphorylated MCM helicase, zones of replication initiation are determined. This allows euchromatin regions to replicate in early S phase, resulting in replication timing. On the other hand, in RIF1-KO cells, the antagonistic phosphorylation regulation of MCM helicase is lost. Because TRESLIN-MTBP can bind to various zones, random replication occurs and replication timing is lost.
Credit: Chikashi Obuse
Future Developments
These results answer the fundamental question of "how human cells initiate genomic DNA replication," and provide new insights into genome instability diseases (such as cell death, cancer, aging, genetic diseases) caused by DNA replication defects, as well as evolution through genome changes. They are also expected to serve as a foundation for the development of new technologies to artificially control DNA replication in the future.
Notes
The article, “Regulated TRESLIN-MTBP loading governs initiation zones and replication timing in human DNA replication,” was published in Nature Communications at DOI: 10.1038/s41467-025-66278-7.
