Multidrug resistance: Not as recent as we thought
Researchers from Osaka University find that a relatively ancient drug efflux pump in Haemophilus influenzae confers resistance to the same drugs as its more evolved counterparts, and is only mitigated by a slightly leaky membrane channel
Researchers from Osaka University have made the striking discovery that multidrug-resistant bacteria may have been around longer than we thought.
In findings published this month in Communications Biology , the researchers investigated the evolutionary relationships among hundreds of RND-type efflux pumps—specialized proteins that pump multiple different types of antibiotics out of a bacterial cell, making it multidrug resistant.
“Interestingly, we found that RND efflux pump AcrB from H. influenzae was relatively ancient but exported the same antibiotics as its more evolved counterpart from Escherichia coli ,” explains lead author of the study Martijn Zwama. “What it couldn’t do well was export bile salts, which are not something that H. influenzae encounters in its normal habitat but are common components of the gut, where E. coli resides.”
While this pointed to evolution of the pumps in their natural environments, it also suggested that multidrug recognition is an ancient trait. This is an important distinction because most bacteria acquire resistance genes or mutations in the face of selective pressure from the environment.
But while AcrB protects H. influenzae from several difference classes of antibiotics, the pathogen remains susceptible to β-lactams and novobiocin, something that researchers have previously not been able to explain.
“Bacterial membranes contain channels that selectively let different substrates into the cell,” says co-author Akihito Yamaguchi. “We found that in H. influenzae , one of these channels, OmpP2, was slightly leaky. This meant that some of the smaller drugs pumped out of the cell by AcrB could seep back inside, where they went to work killing the bacterium.”
Multidrug-resistant bacteria have devastating—and often lethal—effects in infected patients. And with new strains constantly emerging, these super pathogens are arguably the biggest threat to human health today. Therefore, drugs targeting efflux pumps have been developed. However, the researchers found that these drugs had no effect on H. influenzae .
“Efflux pump inhibitors bind to a phenylalanine-rich pocket in AcrB. Unfortunately, these drugs were designed to target the more evolved proteins in species like E. coli ,” explains study corresponding author Professor Kunihiko Nishino. “Because H. influenzae AcrB is more ancient, we found that it does not contain the same pocket configuration and is therefore unaffected by the efflux pump inhibitors.”
Uncovering these evolutionary differences has shown that a “one size fits all” approach is not suitable to address RND-type efflux pumps. This research provides a more accurate picture of the evolution and mechanism of multidrug efflux systems, which will assist the development of new antibiotics to more effectively target specific multidrug-resistant pathogens.
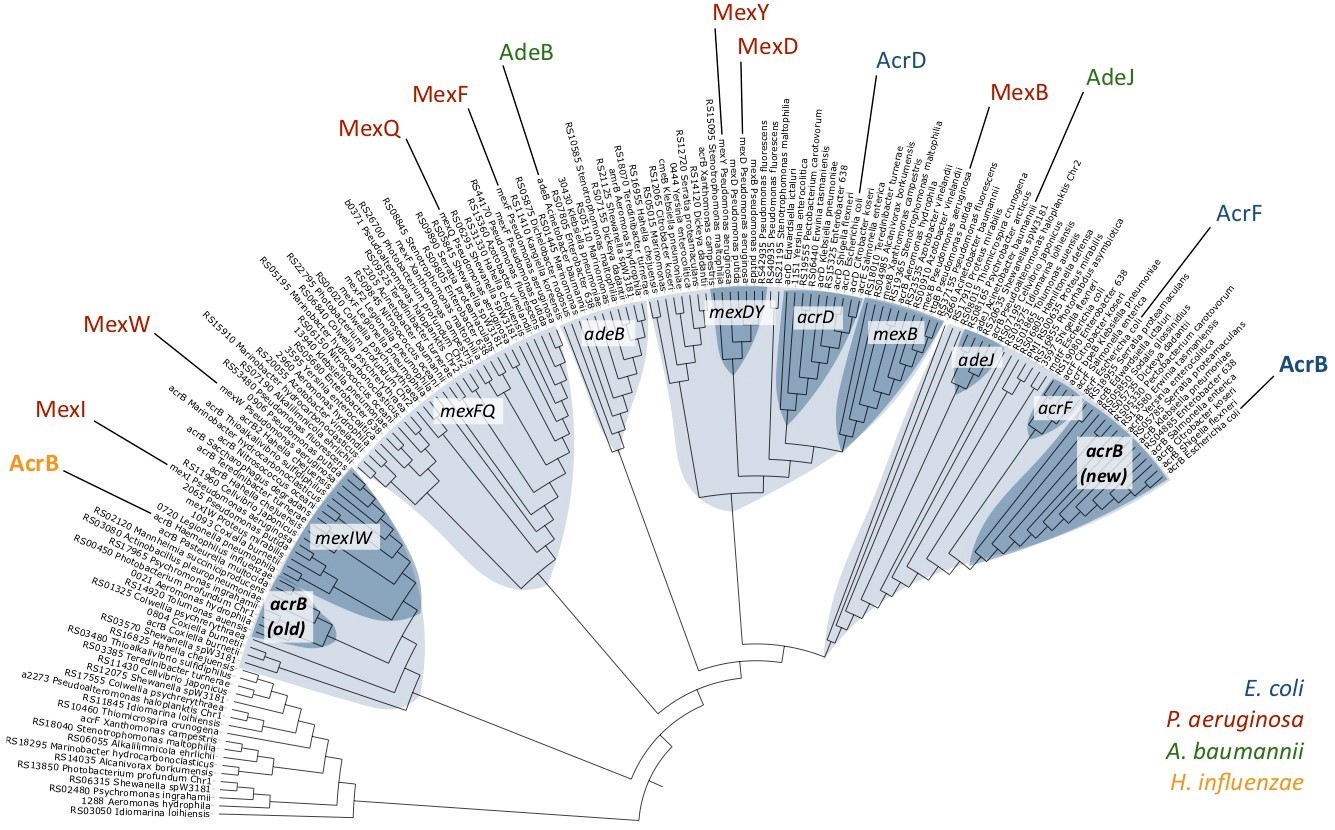
Fig. 1: Phylogenetic relationships among a selection of RND-type MDR transporters. Different clusters are shown in blue, and sub-clusters in dark blue. AcrB-Hi is on the left, and AcrB-Ec on the far right.

Fig. 2: Evolved Phe-rich pit of AcrB-Ec compared with that of the relatively ancient AcrB-Hi.

Fig. 3: Interplay between the efflux pumps and porins from E. coli and H. influenzae . AcrB efflux pumps can actively transport antibiotics out of the periplasm and cytoplasm of bacterial cells, rendering them ineffective. AcrB-Ec can also export bile salts, which are present in the enteric environment where E. coli is found. In comparison, AcrB-Hi showed only weak export of bile salts. Additionally, the efflux pump inhibitor ABI-PP did not affect AcrB-Hi but completely inhibited the efflux ability of AcrB-Ec. The wide OmpP2 outer-membrane porin from H. influenzae leaks small and elongated antibiotics such as β-lactams and novobiocin back into the cells, making Hib cells sensitive to these antibiotics.
The article, “Phylogenetic and functional characterisation of the Haemophilus influenzae multidrug efflux pump AcrB,” was published in Communications Biology at DOI: https://doi.org/10.1038/s42003-019-0564-6 .
Related links
