Synthesis of photosynthetic pigments clarified at a hydrogen atom level
Expected to be applied to the development of devices that convert light into energy
Photosynthetic organisms such as cyanobacterium and plants have a light-harvesting pigment, "bilin." Proteins bound to bilin are involved in photosynthesis in cyanobacterium and the conveying of signals for controlling blooming and defoliating in higher plants. Phycocyanobilin is one of these bilins. This group paid attention to an enzyme, phycocyanobilin:ferredoxin oxidoreductase (PcyA), which helps synthesis of phycocyanobilin by reducing biliverdin. PcyA proceeds with reduction reaction at two different locations. This special reaction is controlled by the atomic position of the enzyme, so examining the structure at the atomic level is important for clarifying how the enzyme proceeds with the reaction.
X-ray crystallography of PcyA has clarified the whole structure of PcyA, interaction between biliberdin and PcyA, as well as amino acids important for synthesizing photosynthetic pigment; however, hydrogen atoms were not seen in the PcyA structure obtained with X-rays. In order to reveal this reaction mechanism, the visualization of hydrogen atoms had been sought after.
In collaboration with Kurume University, Kurume National College of Technology University of Miyazaki, MARUWA Foods and Biosciences Inc., and Ibaraki Prefecture, a group of researchers led by UNNO Masaki (Professor Institute of Applied Beam Science, Graduate School of Science and Engineering, Ibaraki University; Frontier Research Center for Applied Sciences) FUKUYAMA Keiichi (Visiting Researcher, Applied Chemistry, The Graduate School of Engineering, Osaka University; Professor Emeritus, Osaka University), TAMADA Taro (Directorate, Quantum Beam Science, Atomic Science Research Department, Japan Atomic Energy Agency) conducted neutron crystal structure analysis of complex PcyA and its biliverdin, clarifying the structure with a resolution of 1.95Å.
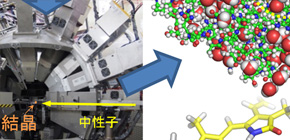
Usually, it's very difficult to visualize hydrogen atoms in protein; however, using the IBARAKI Biological Crystal Diffractometer (iBIX) at the Japan Proton Accelerator Research Complex (J-PARC) in Tokaimura, this group identified the location of hydrogen atoms before reaction of enzyme using hydrogen and found that there were two states: a normal status in biliverdin and a state in which a hydrogen is bound. Furthermore, asparagine, one of the amino acids bound to biliverdin, has two states; a hydrogen-bounded state and hydrogen-not-bounded state. It was also found that hydronium (H 3 O + ) was located between two histidines near biliverdin. These discoveries were obtained because of the visibility of the hydrogen atoms.
In addition, there were differences in results obtained between X-ray crystallography and neutron analysis -- water near biliverdin and interaction between glutamine acid and biliverdin. This suggested that enzyme reaction caused by the reduction ability of the X-rays might have been observed. The most effective means for observing hydrogen atoms is structural analysis using neutrons. This group elucidated the location of hydrogen atoms in the state where PcyA was bound to biliverdin via neutron analysis. This suggested neutron analysis gave more accurate information than X-ray analysis in examining structure of proteins sensitive to reductive reaction.
Abstract
Phycocyanobilin, a light-harvesting and photoreceptor pigment in higher plants, algae, and cyanobacteria, is synthesized from biliverdin IX α (BV) by phycocyanobilin:ferredoxin oxidoreductase (PcyA) via two steps of two-proton-coupled two-electron reduction. We Examined the neutron structure of PcyA from cyanobacteria complexed with BV, revealing the exact location of the hydrogen atoms involved in catalysis. Notably, approximately half of the BV bound to PcyA was BVH + , a state in which all four pyrrole nitrogen atoms were protonated. The protonation states of BV complemented the protonation of adjacent Asp105. The "axial" water molecule that interacts with the neutral pyrrole nitrogen of the A-ring was identified. His88 Nδ was protonated to form a hydrogen bond with the lactam O atom of the BV a-ring. His88 and His74 were linked by hydrogen bonds via H 3 O + . These results imply that Asp105, His88, and the axial water molecule contribute to proton transfer during PcyA catalysis.
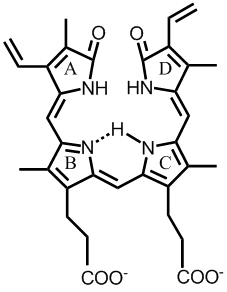
Biliverdin
Figure 1. Phycocyanobilin synthesis reaction catalyzed by PcyA
Figure 2. Success of growing large crystals of PcyA for neutron crystallography
Figure 3. Ibaraki biological crystal diffractometer iBIX
Figure 4. Protonation states of Asp105 and biliverdin
Figure 5. Finding of protonations of His88 and His74, and a hydronium ion
Figure 6. Ripple effects of this research
To learn more about this research, please view the full research report entitled " Insights into the Proton transfer Mechanism of a Bilin Reductase PcyA Following Neutron Crystallography " at this page of the Journal of the American Chemical Society website.
Related Link
