Big Data Provides Clues for Characterizing Immunity in Japanese
An Osaka University-centered research team has performed fine-mapping of the key immunity-related part of the genome, revealing associations between gene variants and over 50 different phenotypes in the Japanese
Although different genes are distributed widely across chromosomes, many genes related to the immune system are clustered together on human chromosome 6, in a segment called the major histocompatibility complex (MHC) region. The density of genes there has made it difficult for researchers to characterize them and their effects, but new technologies and large biobanks with data on huge numbers of people have opened the door to deeper insights into this region.
In a major new study published in the journal Nature Genetics , researchers at Osaka University and their colleagues have surveyed the MHC region specifically in the Japanese population, revealing the existence of different gene variants and their connections with diseases and other traits.
The team based their analyses on three sets of data. One was sequencing information on 33 genes determining white blood cell types in over 1,000 Japanese individuals, obtained by high-throughput sequencing. The second was data from genome-wide association studies looking at links between regions across the whole of the genome and traits and diseases in over 170,000 Japanese individuals. The third set comprised data taken from medical records on over 100 phenotypes reflecting clinical states and other traits.
“Our multiple analyses first revealed the levels of polymorphism in the human leukocyte antigen (HLA) genes, then classified the overall patterns of this polymorphism into 11 distinct groups across the Japanese population using a machine learning approach,” says lead author Jun Hirata. “This provided insight into the genetic landscape of the MHC region and showed us that ‘non-classical’ HLA genes should also be included in efforts to characterize the functional effects of this genomic region.”
After surveying the full complement of variation across the MHC region in the Japanese individuals, the team then focused on clarifying the associations of these variants with different traits and diseases. For this, they used data from medical records on 106 different phenotypes, including 46 complex diseases, from over 170,000 Japanese individuals. About half of these phenotypes showed significant associations with the studied genes. The findings revealed that it is common for a single gene in the MHC region to influence multiple traits, a phenomenon known as “pleiotropy.”
“Our work shows the importance of differences in white blood cell type for health in Japanese people,” senior author Yukinori Okada says. “The cross-phenotype networks that we constructed also showed correlations between health conditions that were not previously known to be related.”
The key findings of this work, including the clinical importance of non-classical HLA genes and the effects of gene variants within MHC haplotypes, should provide a solid foundation for future studies on risk factors associated with this part of the genome.
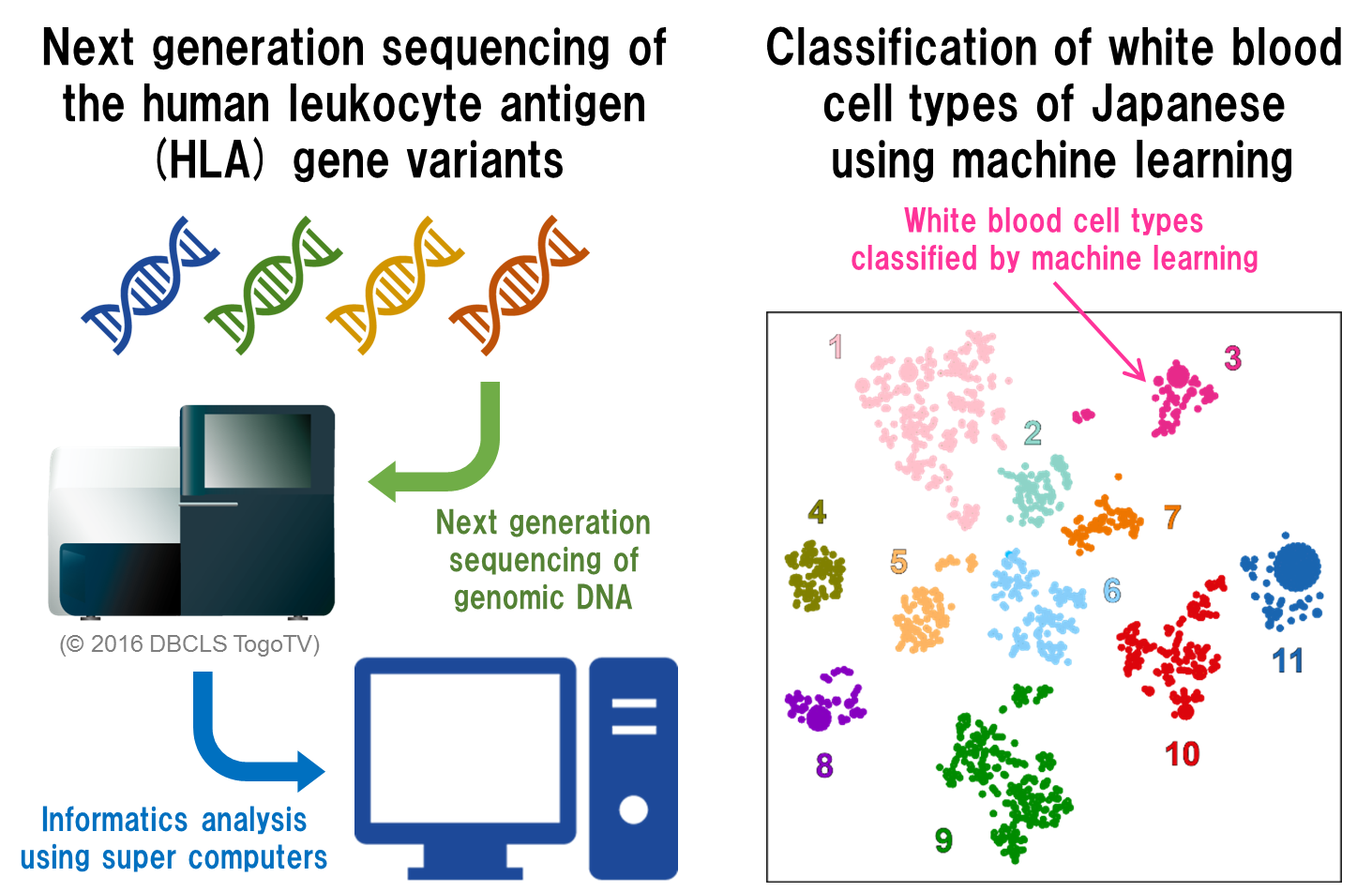
Fig.1 Next generation sequencing of the HLA genes and application of machine learning
(Credit: Osaka University)

Fig.2: The list of the phenotypes associated with the genetic variants in the MHC region
(Credit: Osaka University)
The article “Genetic and phenotypic landscape of the major histocompatibilty complex region in the Japanese population” is published in Nature Genetics at DOI: https://doi.org/10.1038/s41588-018-0336-0 .
Related links
