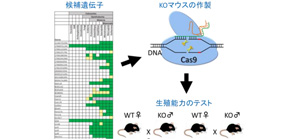
Some genes predominantly expressed in the testis are not related to fertility
Impetus given to an investigation into the cause of infertility via gene analyses using the latest genome editing technology
Knockout mice lacking specific genes of interest are used for examining gene functions at an individual level; however, producing knockout mice cost millions of yen and took 6 to 24 months. The CRISPR/Cas9 system, a new genome editing technology used in this group’s study, has made it possible to produce knockout mice at a cost of hundreds of thousands yen in the short period of time of 1 to 2 months.
This study was conducted using this new genome editing technology. As a result, it has become possible to perform research focusing on genes which have an important role on an individual basis by first investigating their level of importance by using genetically engineered mice and then performing detailed analysis, an approach designed to enhance return on investment.
A group of researchers led by Assistant Professor MIYATA Haruhiko and Assistant Professor FUJIHARA Yoshitaka, and Professor IKAWA Masahito at the Research Institute for Microbial Diseases, Osaka University, in a joint research project with Dr. Julio M. Castaneda, Dr. Zhifeng Yu, and Professor Martin M. Matzuk at Baylor College of Medicine, identified many genes which were expressed in the testis of humans and mice based on theses published in the past and database analysis.
This group produced recombinant mice by using the latest genome editing technology and clarified that 54 genes, or 70 percent of all genes, were not essential for male mice fertility, although they were predominantly expressed in the testis. This group’s achievement has made it possible to perform research focusing on related genes after excluding the 54 genes that are not related to fertility. This approach will significantly improve investment effect of cost and labor of research and lead to the clarification of causes of infertility and the development of vaccines, as well as achieve a remarkable breakthrough in biological research.
Abstract
Gene-expression analysis studies from Schultz et al. estimate that more than 2,300 genes in the mouse genome are expressed predominantly in the male germ line. As of their 2003 publication [Schultz N, Hamra FK, Garbers DL (2003) Proc Natl Acad Sci USA 100(21):12201–12206], the functions of the majority of these testis-enriched genes during spermatogenesis and fertilization were largely unknown. Since the study by Schultz et al., functional analysis of hundreds of reproductive-tract–enriched genes have been performed, but there remain many testis-enriched genes for which their relevance to reproduction remain unexplored or unreported. Historically, a gene knockout is the “gold standard” to determine whether a gene’s function is essential in vivo. Although knockout mice without apparent phenotypes are rarely published, these knockout mouse lines and their phenotypic information need to be shared to prevent redundant experiments. Herein, we used bioinformatic and experimental approaches to uncover mouse testis-enriched genes that are evolutionarily conserved in humans. We then used gene-disruption approaches, including Knockout Mouse Project resources (targeting vectors and mice) and CRISPR/Cas9, to mutate and quickly analyze the fertility of these mutant mice. We discovered that 54 mutant mouse lines were fertile. Thus, despite evolutionary conservation of these genes in vertebrates and in some cases in all eukaryotes, our results indicate that these genes are not individually essential for male mouse fertility. Our phenotypic data are highly relevant in this fiscally tight funding period and postgenomic age when large numbers of genomes are being analyzed for disease association, and will prevent unnecessary expenditures and duplications of effort by others.
To learn more about this research, please view the full research report entitled “ Genome engineering uncovers 54 evolutionarily conserved and testis-enriched genes that are not required for male fertility in mice ” at this page of the PNAS website.
Related links
